Kinetics – or the importance of rate constants in biological and pharmaceutical research
When investigating molecular interactions, equilibrium dissociation constants (KD) are commonly used as a measure of the affinity of two molecules for each other. In other words, the KD is a direct measure of the strength of an interaction. Determining the KD is very important to evaluate the biological relevance of an interaction, such as in the study of fusion proteins, DNA-protein binding and for standard protein analysis.
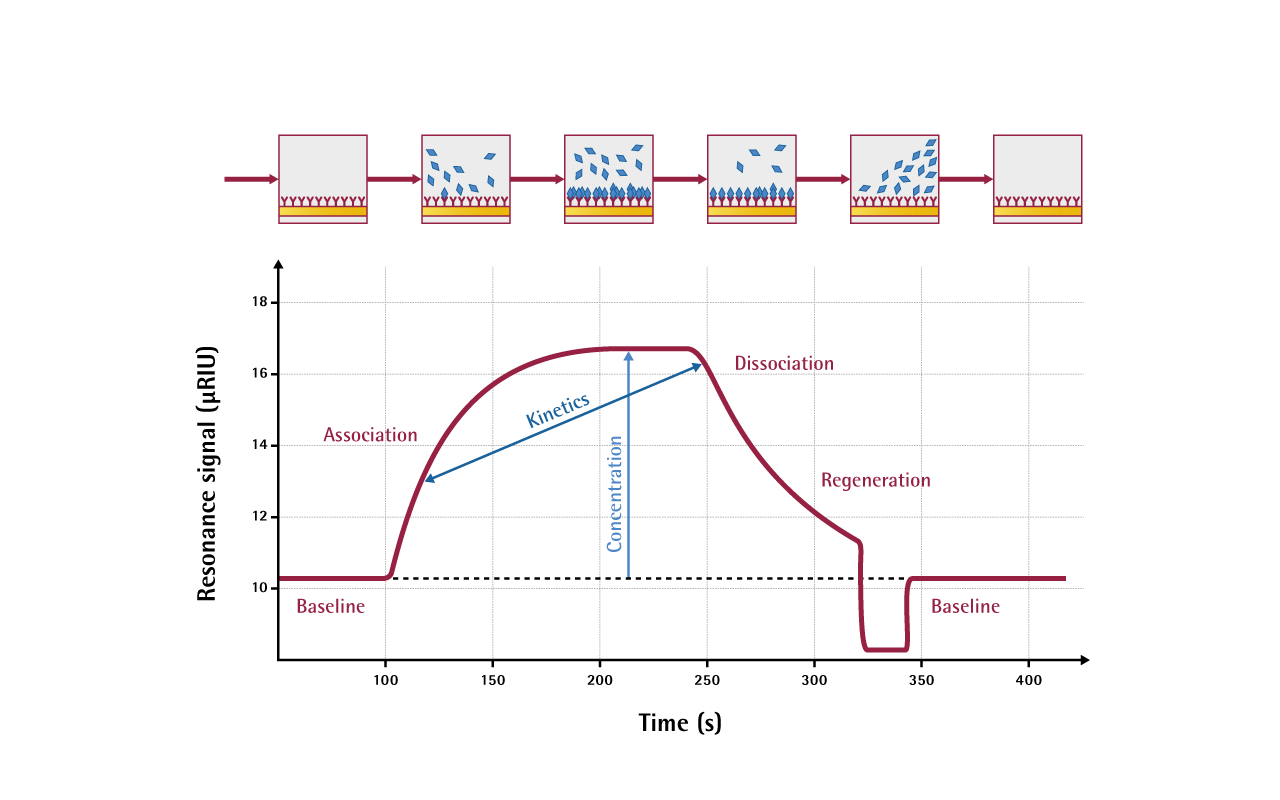
One of the interactants is immobilized on the sensor surface, while the others are free in solution and pass over the surface. A typical cycle of measurement contains association, steady-state and dissociation phases, eventually followed by regeneration; these phases are displayed in the sensorgram.