Surface Plasmon Resonance and Vaccine Research – Developments in Malaria Research
Introduction
Treating a disease as complicated as malaria requires a team approach. Teams of researchers from around the world have been working for many years to determine the best way to combat this deadly disease. Malaria is most prevalent in tropical and subtropical parts of the world including Africa, the Middle East, Southeast Asia and Central and South America1. Development of a vaccine to prevent malarial disease and its transmission are seen as key components of the fight against the disease and its spread. However, finding the proper target for treatment has been complicated, especially since more than one type of parasite can be involved in transmission.
An integral step in malaria vaccine development has been to understand how infection by parasites occurs. Of the known human malaria parasites, the group whose research is highlighted here has targeted two - Plasmodium falciparum (P. falciparum)2, which is the most deadly parasite, and Plasmodium vivax (P. vivax)3 which is the most common parasite species that infects humans. A key question that needed to be answered early on is - How do parasites gain entry into red blood cells? This group discovered that P. vivax enters young red blood cells by making proteins that recognize and bind to receptors on the young red blood cell surface4. This family of proteins was structurally similar to those used by P. falciparum to infect red blood cells. The next step in their research effort was to determine the three- dimensional structure of the proteins. This determination showed that the proteins are folded in the same way with the main difference being in the electrical charge on the surface of the molecules4.
Once researchers worked out the structure of the proteins produced by malaria parasites and where the proteins are binding to their receptors, they started work on designing inhibitors that can potentially block parasite entry. To assess the binding characteristics of these potential inhibitors, researchers turned to surface plasmon resonance (SPR), an information-technique that can determine binding kinetics and affinities for a wide range of biomolecular interactions. This note presents results from a research group that utilized Reichert’s SR7500DC surface plasmon resonance (SPR) system to better define the binding interface of a key malaria protein with its receptor and to show that certain small proteins potentially could be used as soluble inhibitors to inhibit the parasite erythrocyte interaction. This information is an important starting point that will lead to further screening of potentially more powerful inhibitors and can aid in the rational development of a novel malaria vaccine.
Experimental
Background
Researchers found that P. falciparum reticulocyte binding protein-like homologue 4 (PfRh4) binds to complement receptor 1 (CR1 or CD35) to mediate entry of malaria parasites into human red blood cells2. CR1 (complement receptor 1) is composed of 28–30 structural modules (which are called complement control protein (CCP) domains) in the extracellular domain. The CR1 protein fragments in the study outlined here consisted of only two CCP domains and had molecular weights of around 14,000 Daltons. Initial mapping studies identified the first two to three N-terminal modules of CR1 (CCPs 1-2 or CCPs 1–3) as specific inhibitors of the PfRh4-CR1 invasion pathway2. Seven different mutant proteins were tested for their inhibitory potential via SPR. SPR was able to rank them in terms of affinity with three having the highest affinity for the intended target. The SPR results matched with the expected biological activity determined independently with other techniques2.
Conditions
- Instrument: Reichert SR7500DC
- Temperature: 25 °C
- Sensor Chip: XanTec CMD500m
- Running Buffer: 10 mM HEPES pH 7.4, 150 mM NaCl, 1 mM MgCl2, 0.005% tween 20
- Target: Recombinant PfRh4 (P. falciparum reticulocyte binding protein-like homologue 4)
- Analyte: CR1 CCP domains 1-2 (CR1 complement control protein 1-2)
Results
Binding affinities for a series of protein fragments to PfRh4 were determined. An example of responses from one of the higher affinity fragments is show here (CR1 CCP 1-2). Data is fit to a 1:1 binding model with mass transport model:
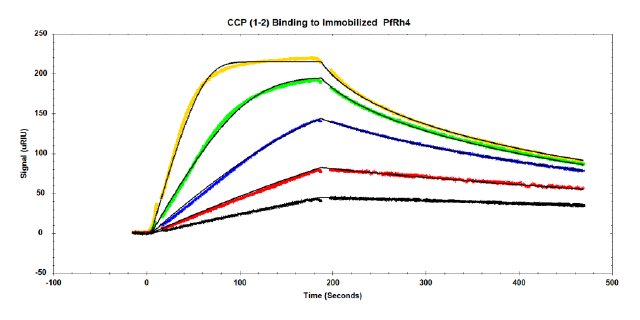
The following concentrations of the CR1 CCP1-2 inhibitor were injected over immobilized PfRh4: 5, 10, 20, 40 and 80 nM. Tracedrawer (Ridgeview Instruments AG) was used to fit the data. The calculated ka is 2.10 e6 M-1 s-1, the kd is 1.26 e-2 s-1, and the equilibrium dissociation constant KD is 6.01 nM.
Summary
- SPR was used to provide important information about the binding interface of a key malaria protein with its receptor.
- SPR results indicate that certain CR1 protein fragments bind tightly and could function as soluble inhibitors to block PfRh4 binding to red blood cells and to inhibit the PfRh4-CR1 invasion pathway2.
- SPR analysis provides critical data on binding affinities that cannot be obtained as readily by other techniques which can aid in efforts to design new vaccines and should be helpful in the ultimate determination of specific single epitope antibodies to block malaria parasite invasion.
References
- CDC website section – Malaria – Malaria Worldwide
- Nicholas T. Y. Lim, Markus J. Harder, Alexander T. Kennedy, Clara S. Lin, Christopher Weir, Alan F. Cowman, Melissa J. Call, Christoph Q. Schmidt and Wai-Hong Tham, Characterization of Inhibitors and Monoclonal Antibodies That Modulate the Interaction between Plasmodium falciparum Adhesin PfRh4 with Its Erythrocyte Receptor Complement Receptor 1,” J. Biol. Chemistry (2015) Vol. 290, pp. 25307-25321, doi:10.1074/jbc.M115.657171.
- Jakub Gruszczyk, Usheer Kanjee, Li-Jin Chan, Sébastien Menant, Benoit Malleret, Nicholas T. Y. Lim, Christoph Q. Schmidt, Yee-Foong Mok, Kai-Min Lin, Richard D. Pearson,,10 Gabriel Rangel, Brian J. Smith, Melissa J. Call, Michael P. Weekes, Michael D. W. Griffin, James M. Murphy, Jonathan Abraham, Kanlaya Sriprawat, Maria J. Menezes, Marcelo U. Ferreira, Bruce Russell, Laurent Renia, Manoj T. Duraisingh, Wai-Hong Tham1, „Transferrin receptor 1 is a reticulocyte-specific receptor for Plasmodium vivax,“ Science (2018), Volume 359, pp. 48-55.
- Walter + Eliza Hall Institute of Medical Research website, Arunee Wilson article on Dr. Wai- Hong Tham, “3D protein map offers new malaria vaccine hope,” updated 19 February 2016.