Capture vs Coupling of an Antibody Using the 2SPR Surface Plasmon Resonance (SPR) System
Introduction
Amine coupling is often the first method a person uses to attach a ligand to an SPR slide. The covalent attachment via free primary amines on a ligand results in random attachment since there are usually multiple attachment sites on the protein. The coupling is done at a pH that is 1-2 units below the pI of the ligand. This facilitates electrostatic attraction to the attachment surface.
A second way to attach a ligand to an SPR slide is via non-covalent capture methods. Non-covalent capture methods are typically used to attach ligands to an SPR surface in instances where the ligand cannot withstand the lower pH needed for covalent coupling or when a ligand needs to be attached in a more oriented manner. This approach is applicable to a variety of types of samples
We provide examples of three different approaches to coupling the monoclonal antibody Anti-Human Serum Albumin (HSA) IgG to an SPR slide in this Application note. The first example is direct amine coupling. The second example is covalently coupling NeutrAvidin to a slide surface and then Capturing biotinylated Anti-HSA. A third example is where Goat Anti-Mouse Fc IgG is amine coupled to a slide surface and then used to capture monoclonal Anti-HSA IgG. In all three cases, antigen binding (HSA) to the Anti-HSA is followed over a series of concentrations and a KD value is determined.
Abstract
Antibody-Antigen binding is an area of interest to many researchers. When deciding to obtain binding kinetics for this type of analysis with Surface Plasmon Resonance (SPR) the question comes up as to what the preferred coupling method is. In this Note we compare three methods of coupling an antibody (monoclonal Anti-HSA IgG) to an SPR slide. The first two methods, direct amine coupling and capture via a biotin tag, both yield stable surfaces where just the analyte (HSA) is removed with regeneration. The third method, capture via a sandwich technique, involves first amine coupling Goat Anti-Mouse Fc IgG to a slide surface, then capturing Anti-HSA, then following the binding of HSA to the antibody. This process involves removal of both Anti-HSA and HSA at each regeneration and so consumes more antibody than the other two techniques. We will explore the plusses and minuses to each of these approaches and compare results obtained with each type of experiment.
Experimental
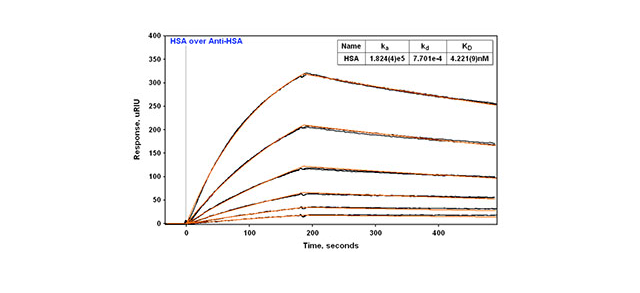
This Application note presents the binding kinetics of a model antibody-antigen system, HSA binding to anti-HSA IgG. The first method involves directly amine coupling the antibody. Terminal carboxyl groups of a planar slide surface are activated using N-Hydroxysuccinimide (NHS) /ethyl(dimethylaminopropyl) carbodiimide (EDC). Primary amines on the antibody are covalently coupled, forming an amide bond. Ethanolamine is injected to cap activated sites not filled by the antibody and to remove loosely bound material. HSA is serially diluted and various concentrations above and below the KD are injected over the surface to obtain binding curves. SPR binding curves obtained are globally fit and a KD value is obtained. A regeneration solution is injected after each sample injection to obtain a fresh surface for the next sample injection (Figure 1A).
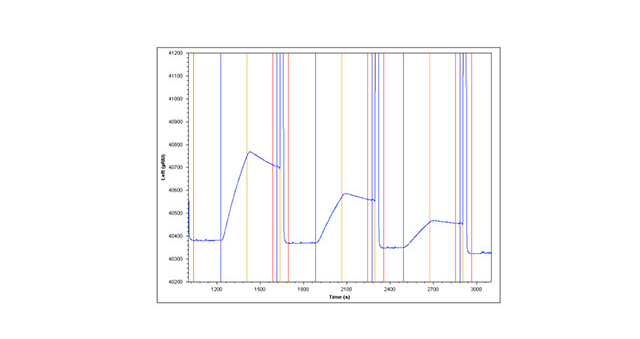
The second method involves capture of Anti-HSA over a NeutrAvidin surface. NeutrAvidin is covalently coupled to the surface of a planar slide with terminal carboxyl groups using standard amine coupling (see method one above for details of the chemistry). Minimally biotinylated Anti-HSA (1.4 moles of biotin per mole of protein) is captured over the planar NeutrAvidin sensor slide creating a stable capture surface. HSA is then injected (as in method one above) to obtain kinetic information about the Anti-HSA/HSA binding interaction (Figure 1A)
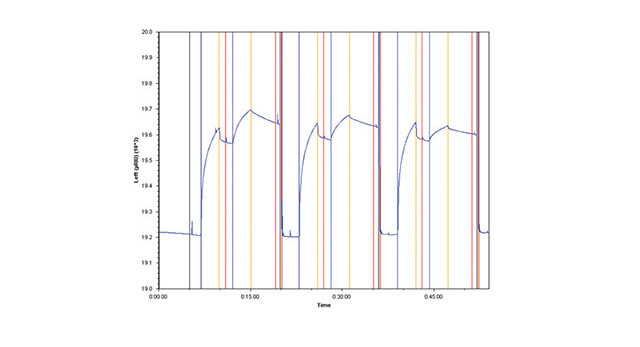
The third method involves amine coupling Goat Anti-Mouse Fc IgG to the sensor chip surface, then capturing Anti-HSA. Each injection of HSA is followed by an injection of regeneration solution that removes both the antibody and antigen (Figure 1B).
Results
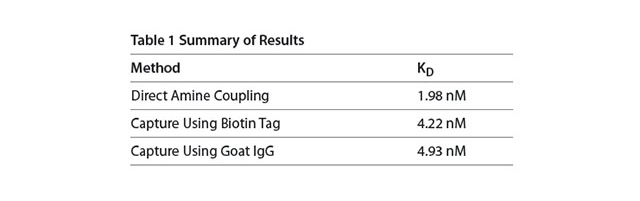
In all three cases the technique is reproducible with replicate injections that overlay with each other (Figure 2). With direct amine coupling, Anti-HSA forms a covalent amide bond to the surface and results have been found to be independent of day and amount coupled (early Application note). A value of about 2 nM for the KD has been determined for this experiment. Comparison to the KD obtained using the two different capture methods is shown in Table 1. The two capture methods yield a very similar KD of 4.2-4.9 nM. One might expect this to be a better reference value since the use of a capture method yields a surface where the antibody is attached in a more specific way (amine coupling is generally considered more random since there are multiple sites for attachment via primary amines on antibodies). In this context, the second method (capture via a biotin tag) is probably most preferred since less antibody is used than with the third method.