There are three methods which are commonly used to couple the bio-specific ligand to the supporting matrix.
The first, and most common, is covalent coupling of the ligand to the matrix. This usually involves cross-reacting functional groups in the dextran layer to amine or thiol groups on the ligand itself. Several different approaches to achieve this can be seen in the following figure.
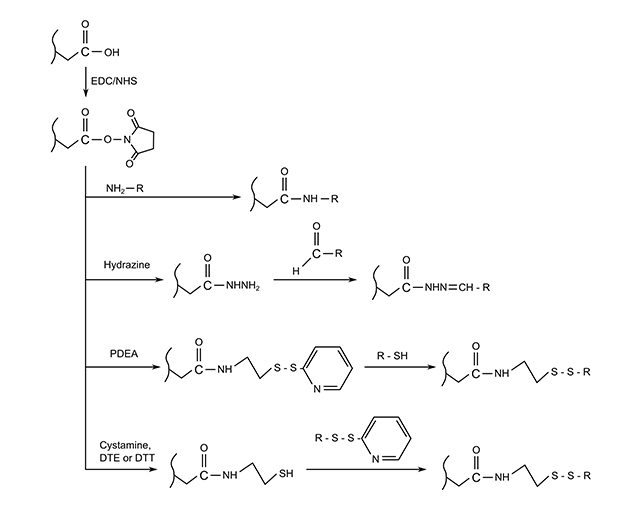
Fig: Four methods for the covalent immobilization of amine, thiol or aldehyde containing ligands on carboxylated surfaces
Secondly, as mentioned previously, biotinylated ligands can be non-covalently linked to streptavidin-coated surfaces. The streptavidin-biotin binding affinity is extremely high, which means that used chips can be regenerated without breaking the non-covalent streptavidin-biotin linkage, (this linkage, however, cannot be regenerated). This approach is well suited for coupling DNA oligomers or PCR products to the sensor surface.
Lastly, one can use capture antibodies. This tactic is usually appropriate when covalent linkage of an antibody has been observed to reduce its activity, and can be thought of as a western blot in reverse. Firstly, a capture antibody (such as mouse anti-Rabbit-Fc) is covalently linked to the surface matrix. Secondly, the antigen-specific antibody (rabbit anti-Tau, for example) is added, where it binds to the capture antibody. Finally the antigen-containing solution is injected, whereupon the antigen will bind to the second antibody. As the majority of these interactions are non-covalent, regeneration of the chip requires that the loading process be repeated.